Laboratory products
Western blotting is a fundamental technique used widely across the life sciences sector by supporting the detection, identification, and quantitation of target protein(s) within a complex sample. Its affordability, specificity and widespread accessibility makes this technique prevalent in protein biology research by offering critical insights into protein expression, localisation and post-translational modifications. However, to obtain reliable western blot results and data integrity requires that researchers must follow best practices for data normalisation. This analytical process facilitates meaningful and accurate comparisons of different samples, minimising variability from experimental errors and sample types [1].
By facilitating precise comparisons of changes in protein expression levels within a given sample set, western blot data normalisation promotes reliable reporting of experimental outcomes, fostering scientific progress and guiding future research [2]. In other words, researchers should be confident that variation observed between target protein abundances and different samples is the result of the actual biology and not due to the data being skewed by common errors, such as inconsistent sample preparation, uneven protein transfer, or pipetting errors.
Data normalisation using loading controls is essential for unbiased western blot experiments, eliminating process errors that could lead to variations in sample amounts across lanes. Although experiments should aim to minimise such inconsistencies through proper design and execution, data normalisation corrects for any non-biological differences between test samples and target proteins prior to comparison.
Despite the different approaches available for researchers to normalise their data, there has been a growing crisis of confidence within the scientific community that has led many scientific journals to increase the stringency of their publication guidelines [3]. Researchers should thoroughly evaluate available normalisation methods and associated guidelines surrounding their use. Regardless of the chosen technique, achieving reliable results hinges on establishing a linear detection range in data normalisation, to demonstrate that any protein signal changes align proportionally with the amount of protein contained within test samples run on the same blot [1].
There are two main types of normalisation that can be applied when performing western blots: single protein detection normalisation and total protein normalisation (TPN) - each with its own set of advantages and potential drawbacks for researchers to consider.
Single protein detection uses proteins, such as ‘housekeeping’ proteins (HKPs) (i.e., internal normalisation standards) or exogenous proteins as loading controls [4]. Immunoprobing these proteins determines their abundance, serving as a proxy for the entire protein population within a sample.
To qualify as a loading control, an internal protein must meet specific criteria: it cannot be your target protein and must exhibit abundant, ubiquitous, and consistent expression within the sample. HKPs, such as β-actin, GAPDH, and β-tubulin, are commonly chosen as internal normalisation standards due to their involvement in basic cellular functions and widespread expression. Consequently, antibodies for them are widely accessible, and their expression is easily detectable.
To ensure the reliability of housekeeping protein (HKP) expression, it is crucial to validate positive and negative controls experimentally, confirming consistent expression levels across sample populations, and establishing the linear range of detection [5]. This latter step is essential because while HKPs are typically abundantly expressed, target proteins often exhibit low expression levels. Overloading samples to detect low-abundance target proteins can saturate HKP reference bands beyond the linear range, undermining their function as loading controls and potentially leading to inaccurate detection.
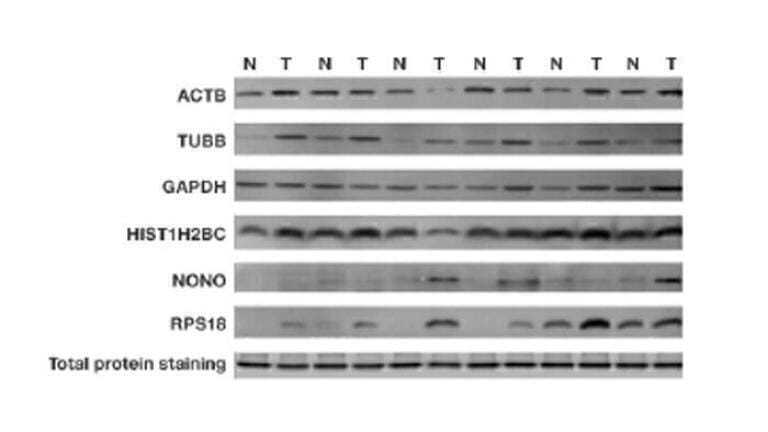
Despite their ubiquitous expression, recent studies have also shown that the expression levels of HKPs themselves can vary across different cell types and are influenced by different biological factors, including the cell cycle, cell density, developmental cycle, tissue type, subject age, and response to treatment [6]. In addition, HKP expression has been shown to be upregulated in certain cancers (Figure 1) [7]. As a result, it is important for researchers to validate their HKP for consistent expression across different sample types and experimental conditions. This can be achieved by conducting a linear range determination test, which may involve several rounds of primary and secondary antibody dilution ratio optimisation.
Figure 1: Differences between HKP expression and total protein staining between tumour (T) and non-tumour (N) cancerous tissues. Taken from Hu et al [7].
Exogenous protein normalisation is a less common technique for single protein detection, whereby a known quantity of purified or recombinant protein is added into the samples. Given that the primary antibody is capable of solely detecting this exogenous protein, this type of control offers the ability to precisely control the amount of protein added and ensures consistency across all experimental conditions. Whilst control tests are also necessary for exogenous proteins, they bypass the need for an expression level test.
When using an exogenous control, however, it is important to take the point at which the protein is introduced into consideration. For example, when the protein is introduced after sample extraction from cells or tissues, it can only be used to normalise experimental error introduced through sample loading and gel-to-membrane transfer and not for variation in sample preparation.
TPN provides an alternative approach whereby all sample proteins are visualised, and total protein levels are quantified. This eliminates the reliance on a single control protein to represent the entire protein population. Instead, TPN utilises the total protein abundance measured from the blot as the basis for normalisation.
Colorimetric dyes such as Ponceau S, Coomassie Blue and Amido Black can be measured via densitometry or imaged with a white light source of a digital imager, whilst stains such as Sypro RUBY can be measured via a fluorescent light source from a digital imager. As Ponceau S and Sypro RUBY are reversible dyes, staining can be performed prior to immunodetection, but staining and destaining steps are time sensitive so researchers must perform these steps with care. On the other hand, Coomassie Blue and Amido Black are irreversible stains and can therefore be used after immunodetection steps [2].
Total protein stains are typically less sensitive than antibody-based immunodetection used for detection of HKPs, overcoming issues related to oversaturation and therefore demonstrate good linearity within a typical loading range of 10–50 μg of cell lysate enabling a more accurate determination of low-abundance target proteins. In addition, normalising target protein levels against total protein expression prevents results being affected by variable HKP expression (Figure 1). TPN is also less likely to be impacted by variability in the reference signal, which is more likely to occur using HKP normalisation due to the need for stripping and reprobing steps. Consequently, despite the historical precedence of HKP normalisation, there is increasing advocacy for TPN as the method of choice for normalisation in western blotting.
As with HKP normalisation, experiments to validate the positive and negative controls, and expression level test and linear range determination control experiments are required. However, once a sample dilution series has been established, running a single blot and gel can serve as the basis for all control experiments and is not impacted by the behaviour of a particular protein, unlike normalisation with HKPs which requires a new set of control experiments to be performed to optimise the signal or when experimental conditions change.
TPN can also be conducted using stain-free methods, which were introduced to overcome issues related to different membranes having different sensitivities to the stains used in TPN and provides a method to visualise the gel and blot with an imaging process that does not interfere with downstream immunodetection steps [8]. Stain-free technology also avoids the need for additional staining steps that would otherwise be required to visualise strong total protein signals with low background (Figure 2).
Figure 2: Comparison between high background and low background signals in gels with (left) and without (right) Stain-Free technology. Taken from Oh and Turner [1].
Visualisation of total protein on a gel or membrane in stain-free TPN is enabled through a proprietary trihalo compound that is directly embedded into the polyacrylamide gel. Following UV activation, the compound enhances natural protein fluorescence by covalently binding to tryptophan residues, enabling visualisation of total protein without the need for staining and destaining steps (Figure 3).
Figure 3: Exposure to UV light activates the embedded Stain-Free compound within Stain-Free gels, enabling visualisation of total protein. Taken from Wang [8].
In comparison to HKP normalisation, the overall workflow for stain-free TPN is considerably simplified as running a single gel and blot serves as the basis for all control experiments. It also does not require any additional buffers or reagents, making it compatible with existing western blot workflows. Furthermore, stain-free TPN eliminates the issues related to the use of HKPs as loading controls as it is far more resistant to signal saturation within the common loading range for cell lysates. As a result, stain-free TPN establishes a linear and scalar response, allowing researchers to accurately quantify both the target protein and loading control within a linear dynamic range by normalising bands to total protein in each lane (Figure 4). In contrast, a scalar response is not observed with HKP expression, due in large part by combining chemiluminescence – a very sensitive signalling technique - with the immunodetection of a high abundance target making it challenging to differentiate any differences in lysate abundance. Correcting this issue often involves additional control experiments to optimise HKP antibody dilution ratios or using fluorescent HKP antibodies with a wider linear dynamic range and signal response, which increases experimental effort and duration.
Figure 4: Linearity comparison of Stain-Free total protein measurement and immunodetection of three housekeeping proteins in 10-50 μg of HeLa cell lysate. Adapted from Hammond et al [9].
In line with recent discussions regarding the accuracy of normalisation in western blotting, several scientific journals, including the Journal of Biological Chemistry, have since updated their submission guidelines [10]. These show an increasing preference for TPN over HKP normalisation methods and reinforce the importance of transparency and reproducibility when reporting results from experimental data. Consequently, it is even more important for researchers to carefully consider their choice of normalisation method. Adhering to updated journal guidelines whilst maintaining the highest standards of experimental best practice and transparency when reporting data will increase researchers’ confidence in their results, providing clearer insights into their investigations.
1. Oh K & Turner L (2018) The how and why of normalizing your western blots, https://www.bioradiations.com/the-how-and-why-of-normalizing-your-western-blots/
2. Bio-Rad, Total Protein Normalization. https://www.bio-rad.com/en-uk/applications-technologies/total-protein-normalization?ID=PODYJQRT8IG9
3. Fosang A J & Colbran R J (2015) Transparency Is the Key to Quality. The Journal of biological chemistry, 290(50), 29692–29694. https://doi.org/10.1074/jbc.E115.000002
4. Oh, Western Blot Normalization Methods. https://info.bio-rad.com/western-blotting-webinar-LP1.html
5. Janes K A (2015) An analysis of critical factors for quantitative immunoblotting. Science Signaling, 8(371), rs2. https://doi.org/10.1126/scisignal.2005966
6. Nie X et al (2017) An appropriate loading control for western blot analysis in animal models of myocardial ischemic infarction, Biochemistry and biophysics reports, 12, 108-113. https://doi.org/10.1016/j.bbrep.2017.09.001
7. Hu X et al (2016) Common housekeeping proteins are upregulated in colorectal adenocarcinoma and hepatocellular carcinoma, making the total protein a better “housekeeper”. Oncotarget, 7(41), 66679–66688. https://doi.org/10.18632/oncotarget.11439
8. Wang (2021) Stain-Free Western Blotting: Faster Results, Better Data, https://www.bioradiations.com/stain-free-western-blotting-faster-results-better-data/
9. Hammond et al (2020) A Method for Greater Reliability in Western Blot Loading Controls: Stain-Free Total Protein Quantitation, https://www.bio-rad.com/webroot/web/pdf/lsr/literature/Bulletin_6360.pdf
10. Collecting and Presenting Data, Journal of Biology Chemistry. https://jbcresources.asbmb.org/collecting-and-presenting-data
Katie Schaefer earned a PhD in Biochemistry studying gene regulation by transcription factor p53 in response to DNA oxidative stress. She began her career as a field applications scientist with Bio-Rad Laboratories before moving into Global Marketing for chromatography and digital imaging products.
Kenneth Oh earned his PhD in bioorganic chemistry at UC Santa Barbara, where he synthesised fluorescence-based peptide beacons that measured nucleic acid and protein binding events. On graduating, he cofounded a biotech company that reengineered virus capsids to serve as a drug delivery platform. He is currently the Marketing Manager for the Protein Quantitation Business Group at Bio-Rad Laboratories, responsible for the Electrophoresis and Western Blotting imagers, consumables, and reagents portfolio.
Deanna Woo is currently a Global Product Manager for Bio-Rad Laboratories’ Protein Quantitation Global Business division. She received her Bachelor of Science at San Francisco State University in Cellular and Molecular Biology, and held research roles at UCSF, Genentech, and Stanford. Her previous research work included studying the molecular mechanisms that regulate the development of primordial germ cells and research exploring hyperphosphorylation of tau in neurodegenerative diseases.
Lab Asia 33.2 April


