-
-
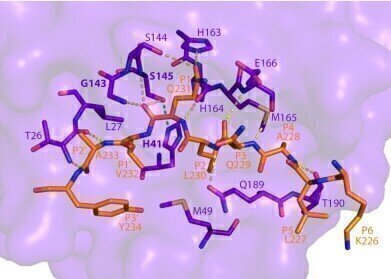 This image shows how SARS-CoV-2 Mpro recognizes and cuts NEMO based on the crystal structure determined using a powerful X-ray beam at SSRL Beam Line 12-2. (Credit : SLAC National Accelerator Laboratory)
This image shows how SARS-CoV-2 Mpro recognizes and cuts NEMO based on the crystal structure determined using a powerful X-ray beam at SSRL Beam Line 12-2. (Credit : SLAC National Accelerator Laboratory)
News & Views
Critical Interaction reveals strength of SARS-CoV-2 Protein
Oct 06 2022
Scientists at the Department of Energy’s SLAC National Accelerator Laboratory recently witnessed the moment when a SARS-CoV-2 virus protein Mpro cut a protective protein known as NEMO, in an infected person. Without NEMO, an immune system is slower to respond to increasing viral loads or new infections.
Funnelling powerful x-rays from SLAC’s Stanford Synchrotron Radiation Lightsource (SSRL) onto crystallised samples of the protein complex, revealed how Mpro attacks NEMO at the molecular level as it dismantled NEMO’s primary function of helping our immune system communicate.
“We saw that the virus protein cuts through NEMO as easily as sharp scissors through thin paper,” said co-senior author Soichi Wakatsuki, professor at SLAC and Stanford. “Imagine the bad things that happen when good proteins in our bodies start getting cut into pieces.”
The images from SSRL showed the exact location of NEMO’s cut and provided the first structure of SARS-CoV-2 Mpro bound to a human protein.
“If you can block the sites where Mpro binds to NEMO, you can stop this cut from happening over and over,” SSRL lead scientist and co-author Irimpan Mathews said. “Stopping Mpro could slow down how fast the virus takes over a body. Solving the crystal structure revealed Mpro’s binding sites and was one of the first steps to stopping the protein.”
NEMO is a critical part of the human immune systems protective inflammatory response; when cut it helps the virus evade innate immune responses. A separate study by researchers at institutions in Germany found that the loss of NEMO by the action of Mpro could lead to damage in certain brain cells, causing neurological symptoms observed in COVID-19 patients, the researchers said.
“The crystal structures of NEMO and Mpro provide us with the targets to develop treatments that stop these cuts from happening,” SLAC scientist and co-first author Mikhail Ali Hameedi said. “Although current antiviral drugs can target Mpro, seeing the molecular details of how Mpro attacks NEMO will help us develop new treatments in the future as Mpro mutates.”
To predict how well Mpro binds to NEMO, researchers used the Summit supercomputer at the Oak Ridge Leadership Computing Facility, combining molecular dynamics simulations with five machine learning models. Applying quantum chemistry, they found that Mpro likely has the highest binding affinity in SARS-CoV-2 compared to the other primary coronaviruses.
“With a set of computational approaches, we were able to predict the strongest binding spots between NEMO and Mpro,” co-first author and ORNL scientist Erica Prates said. “We think that a high binding affinity at these hot spots helps explain the high fitness of the virus in humans.”
Moving forward, the biomedical industry could use the study to help build better inhibitor drugs and understand how other proteins could be affected by Mpro, Wakatsuki said.
“NEMO is only the tip of the iceberg,” he added.. “We can now study what happens when many other proteins in the body are cleaved by Mpro during infection.”
More information online
Citation: M. A. Hameedi et al., Nature Communications, 8 September 2022 (10.1038/s41467-022-32922-9)
Digital Edition
ILM 49.5 July
July 2024
Chromatography Articles - Understanding PFAS: Analysis and Implications Mass Spectrometry & Spectroscopy Articles - MS detection of Alzheimer’s blood-based biomarkers LIMS - Essent...
View all digital editions
Events
Jul 28 2024 San Diego, CA USA
Jul 30 2024 Jakarta, Indonesia
Jul 31 2024 Chengdu, China
ACS National Meeting - Fall 2024
Aug 18 2024 Denver, CO, USA
Aug 25 2024 Copenhagen, Denmark

.jpg)

24_06.jpg)













