-
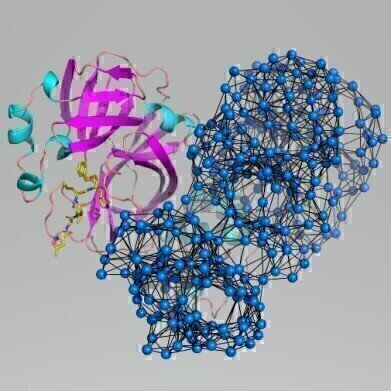 Scientists have used computer modelling to identify potential ‘vulnerable sites’ on a key protein found in coronavirus (Credit: University of York)
Scientists have used computer modelling to identify potential ‘vulnerable sites’ on a key protein found in coronavirus (Credit: University of York)
News & Views
Modelling identifies Vulnerable Sites on Coronavirus Protein
Jan 16 2021
University of York scientists have been able to identify potentially vulnerable sites on a key protein found in coronavirus using computer modelling, paving the way for possible new drug treatments in the future.
The coronavirus responsible for the Covid-19 epidemic deploys dozens of viral biomolecules when it invades host cells with the disease. One of these is a compact protein, the main protease, whose function is critical to the virus. By analysing the structure of the protease using modelling techniques, the scientists have been able to simulate the protein’s motions, suggesting sites that might be accessible to new drugs.
The study(1), by Tom McLeish, Professor of Natural Philosophy in the Department of Physics and Igors Dubanevics from the School of Natural Sciences, ‘was not related to the current vaccines, which are based on the ’Spike’ protein, but is a study of another key protein in the Covid process,’ Prof McLeish said.
He added: “It is more relevant to potential future drugs than to future vaccines, as the motions of the protein that it uncovers point to new ’sites’ on the protein where binding small molecules might disrupt the protein function. The advantage of these sites and our method in general, is that they are not the ‘obvious’ ones that compete with the normal binding of the protein, but other sites that can be accessed even when the usual binding sites are occupied.”
Igors Dubanevics said: “We have identified promising druggable sites in the main protease via computer simulations and some of them have been supported by the newest studies by other groups. The next logical step would be to investigate the identified sites by conducting biological experiments in a lab.
(1) Published in the Journal of the Royal Society Interface.
Digital Edition
Lab Asia 31.2 April 2024
April 2024
In This Edition Chromatography Articles - Approaches to troubleshooting an SPE method for the analysis of oligonucleotides (pt i) - High-precision liquid flow processes demand full fluidic c...
View all digital editions
Events
May 05 2024 Seville, Spain
InformEx Zone at CPhl North America
May 07 2024 Pennsylvania, PA, USA
May 14 2024 Oklahoma City, OK, USA
May 15 2024 Birmingham, UK
May 21 2024 Lagos, Nigeria










.jpg)